A Level Biology - using the flexibility of OCR’s PAGs
01 October 2021
 Jon Hale, Head of Biology at Beaulieu Convent School in Jersey
Jon Hale, Head of Biology at Beaulieu Convent School in Jersey
At our centre we try to incorporate practical activities in the heart of the specification. We found that OCR’s flexible approach to practical endorsement allowed us to adjust our teaching according to our students’ needs. In this blog I share my experience of teaching practical endorsement and how I adapted PAG10 to meet the resources available in our centre.
Our approach to practical endorsement
We do not manage the practical endorsement as a separate entity to the content, because this can become a high stakes venture for our students. We try to provide as many opportunities as possible for students to develop competency for each skill, by ensuring that every learner can meet the criteria over the two years without the anxiety.
This means that each of our students will typically undertake over 60 practical activities, which is where the flexibility of the OCR specification comes into its own.
Also, we do not assess every skill with every practical within the relevant group. We use some of OCR’s suggested practical activities, and sometimes adjust them or we use our own.
The flexible PAG tracker allows us to use our own practical activities and map the skills quickly.
PAG 10 was one of the first practicals we had to adjust to resources available in our centre. Our students are allowed to bring their own devices at school with most of them using Chromebooks. This has made the computational modelling of DNA using RasMol exceptionally challenging over the past few years.
Adaptations to PAG 10 activities
At this year’s ASE Conference, I was introduced to the browser based Mol*, which works with intuitive controls for tablets and Chromebooks. This ease of access moves away from the command-line controls in RasMol, but it maintains the authenticity as it is frequently used by scientists over other molecular viewers. Although, this has made it possible for students to meet all the criteria on their own devices, in school or at home, it needed some adaptations from the original OCR PAG10 suggested activities.
Students can follow the written instructions with relative ease following a short tutorial on some of the nuances of the viewer, such as where the selection cursor is and how to screenshot. This enables students to focus their attention on the molecules rather than “discovery-based learning” of the controls.
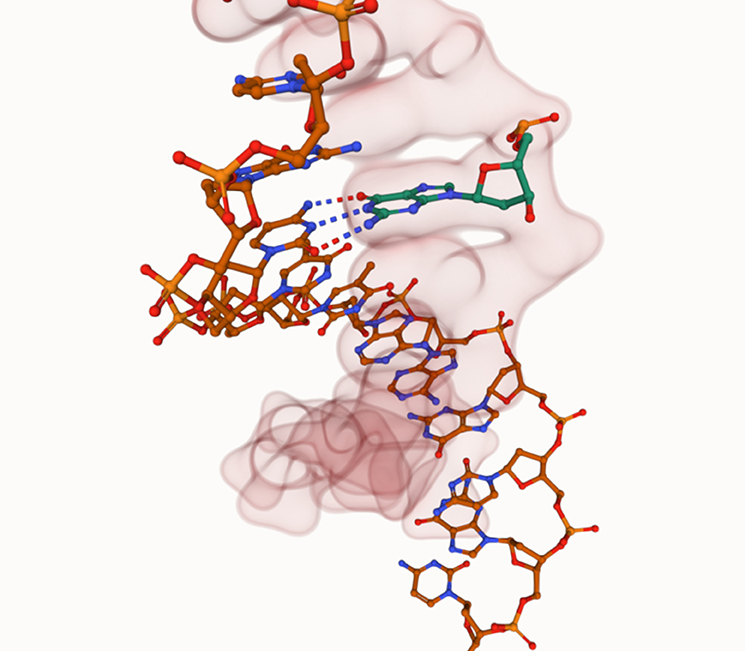
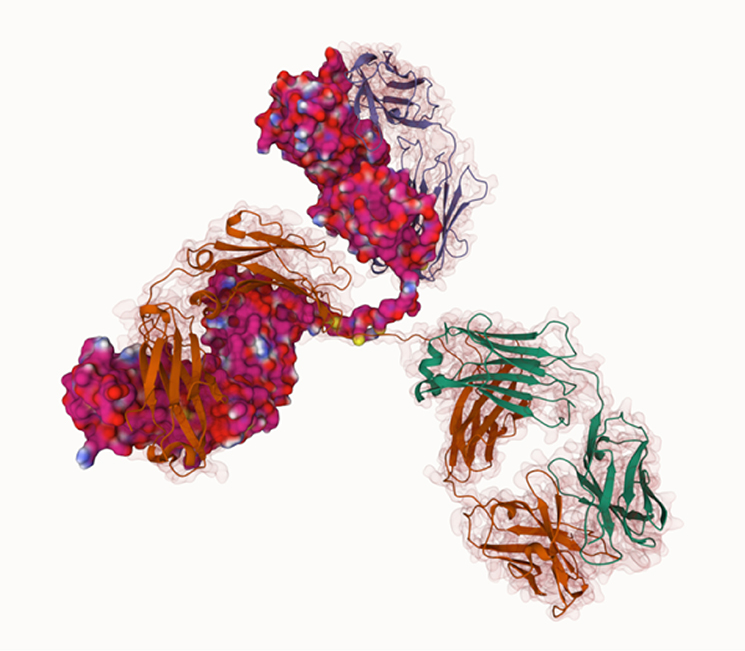
This direct replacement of a PAG10 is not the only way we have used Mol*. As mentioned, our ethos is in providing frequent embedded practicals in the context of the content rather than a bolt-on. So we have Mol* practicals for catalase (enzymes), antibodies (immunity) and RuBisCO (photosynthesis).
The regular application of computational modelling in visualising biological molecules is aiding students’ recognition of globular shapes, diversity, and co-factors along with improving their understanding of the level of protein structure.
Going beyond
In a slightly different approach, we developed a PAG10 activity based on data from the Human Genome Project within the teaching of gene technologies. In this activity students design primers and a thermocycler programme to amplify their region of interest. This then feeds into a PAG6 activity with the students PCR-ing their own DNA before running a gel electrophoresis analysis.
Sequences like this help show the connections between different practical activity groups. It also provides opportunities for students to consolidate their conceptual understanding.
The support provided by the subject advisors at OCR over the past few years has been beneficial in developing these activities. The next quest will be to see if we can use AlphaFold in the classroom!
Stay connected
If you have any comments or questions, you can email us at science@ocr.org.uk, call on 01223 553998 or tweet us @OCR_Science. You can also sign up to subject updates to receive information about resources and support.
About the author
Jon Hale is Head of Biology at Beaulieu Convent School in Jersey. He has been teaching for 15 years and has recently enrolled on a part-time PhD at the University of Dundee. He also represents the interests of school teachers on the committee of the British Ecological Society’s Teaching and Learning Special Interest Group.